Leave message
Can’t find what you’re looking for?
Fill out this form to inquire about our custom protein services!
Inquire about our Custom Services >>






























 Order Online! Now! Get your $50 coupon for online order.
Order Online! Now! Get your $50 coupon for online order. Order Online! Now! Get your $50 coupon for online order.
Order Online! Now! Get your $50 coupon for online order.
 Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!  Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!
 Fill out organ-on-a-chip questionnaire to win a FREE gift!
Fill out organ-on-a-chip questionnaire to win a FREE gift!  Fill out organ-on-a-chip questionnaire to win a FREE gift!
Fill out organ-on-a-chip questionnaire to win a FREE gift!
> Insights > Main characteristics of “Deltacron” according to three cases in France The Pasteur Institute has recently published a preprint paper in In medRxiv entitled "Culture and identification of a "Deltamicron" SARS-CoV-2 variant in a three cases cluster in southern France. Maria Van Kerkhove, the head of COVID-19 technology at the WHO, said at a press conference that she was concerned about Deltacron and confirmed that the strain had begun to appear in France, Denmark, Germany and the Netherlands.
According to PCR results, the scientists confirmed that cases in France were caused by a recombinant strain of Delta 21J/AY.4 and Omicron 21K/BA.1, which was named as "Deltacron". In this SARS-CoV-2 recombinant, most of the spike gene was replaced in a Delta 21J/AY.4 matrix by an Omicron 21K/BA.1 sequence.
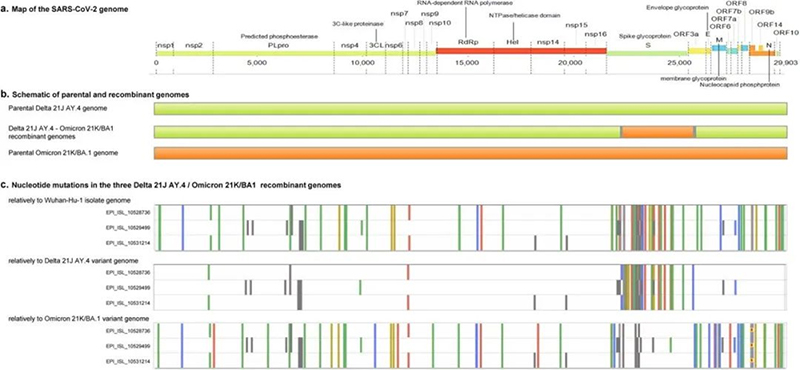
Figure 1. Schematic of the SARS-CoV-2 Delta 21J/AY.4-Omicron 21K/BA.1 recombinant genome.
The overall structure of the recombinant spike protein has been predicted according to the sequence of the recombinant virus. Compared to the Delta and Omicron variant strains, the main structural changes of this recombinant strain were located in the N-terminal domain (NTD), which is attracted by lipid rafts and provides electronegative landing platforms for the spike in the initial interaction of the virus. In this region, the surface of the recombinant spike protein is enlarged, flattened, and more electropositive. Therefore, an increase in the electrostatic surface of the NTD is expected to accelerate the binding of the virus to lipid rafts, which may confer a selective kinetic advantage against virus competitors. The receptor-binding domain (RBD) of the AY.4 x BA.1 recombinant strain is mainly inherited from the Omicron variant strain. The recombination leads to an increase in the electrostatic surface potential of the RBD, which may facilitate the interaction with the electronegative interface of the ACE2 cellular receptor. Overall, this structural analysis suggests an optimization of virus binding to the host cell membrane of the recombinant virus.
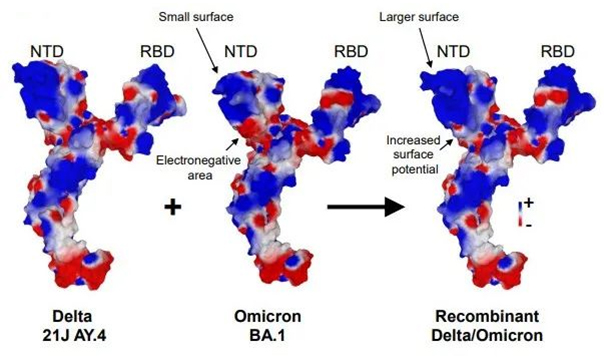
Figure 2. Schematic of the predicted structure of the spike protein of the SARS-CoV-2 Delta 21J/AY.4-Omicron 21K/BA.1 recombinant
With the authenticity of the AY.4 x BA.1 recombinant variant strain being further confirmed, the key issues that need to be considered are again aroused:
- Is it easier to replicate and spread?
- Does it cause more hospitalization?
- Can the existing diagnostic reagents detect it?
- Are currently used therapeutic drugs still effective?
- Is it easier to escape before vaccination or natural immunity?
In order to support the quick response to Deltacron, ACROBiosystems has developed a series of antigens of the recombinant virus for the first time according to the GISAD database, including Spike trimer, S1 and NTD. The product would be strictly controlled to ensure high quality and bioactivity. You can preorder any of these products to make sure you are able to get them as soon as possible.
Product list
Click on the image to view product details
| Molecule | Cat.No. | Tag | Host | Mutation |
|---|---|---|---|---|
| Spike protein | SPN-C5226 | His Tag | HEK293 | T19R, A27S, T95I, G142D, EFR156G, NL211I, INS214EPE, G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y, Y505H, T547K, D614G, H655Y, N679K, P681H, N764K, D796Y, N856K, Q954H, N969K, L981F, R683A, R685A, F817P, A892P, A899P, A942P, K986P, V987P |
| Spike S1 | S1N-C52Hw | His Tag | HEK293 | T19R, A27S, T95I, G142D, EFR156G, NL211I, INS214EPE, G339D, S371L, S373P, S375F, K417N, N440K, G446S, S477N, T478K, E484A, Q493R, G496S, Q498R, N501Y, Y505H, T547K, D614G, H655Y, N679K, P681H |
| Spike NTD | SPD-C522m | His Tag | HEK293 | T19R, A27S, T95I, G142D, EFR156-158G, N211del, L212I, ins214EPE |
>>> click to find out more products about other SARS-CoV-2 mutants .
This web search service is supported by Google Inc.







