 Request a FREE Sample of our FcRn Binding Kit!
Request a FREE Sample of our FcRn Binding Kit! Request a FREE Sample of our FcRn Binding Kit!
Request a FREE Sample of our FcRn Binding Kit!
 Limited Edition Golden Llama is here! Check out how you can get one.
Limited Edition Golden Llama is here! Check out how you can get one.  Limited Edition Golden Llama is here! Check out how you can get one.
Limited Edition Golden Llama is here! Check out how you can get one.
 Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!  Request a FREE sample of our GMP products!
Request a FREE sample of our GMP products!
> Insights > Omicron recombinants and new variants: Who is the next master for immune escape? Global epidemic update
- According to Worldometer, the total number of confirmed COVID-19 cases exceeded 520 million on May 15, 2022 with about 480,000 new cases worldwide within a single day.
- he latest statistics from Johns Hopkins University shows that the cumulative number of COVID-19 deaths in the United States has exceeded 1 million as of May 17, 2022, with 8,260,909 confirmed cases.
- According to a WHO assessment released on May 5, about 14.9 million "excess deaths" are caused by COVID-19 from January 1, 2020 to December 31, 2021.
The risk of the pandemic still remains. The number of COVID-19 cases is rising again in America and Europe. The continuous mutation of Omicron causes a surge in cases in South Africa and other countries. "The COVID-19 pandemic is only entering a new phase and is far from over," the European Commissioner said.
In March 2022, the confirmation of the novel COVID-19 recombinant strain "Deltacron" (XD) breaks out in the news. The World Health Organization (WHO) considers that the risk of COVID-19 evolution and the emergence of new and mutated strains, including recombinant strains, remains high. since the discovery of XD, various recombinant strains such as XE and XL have appeared. Until now, a total of 21 recombinant strains from XA to XW (skipping XI and XO) have been recorded in the Pangolin database. researchers say that the recombination of variants is actually not uncommon, which may occur when the body is infected with two or more COVID-19 variants at the same time. Although the vast majority of recombinant variants die out quickly there is opportunity for more transmissible strains.
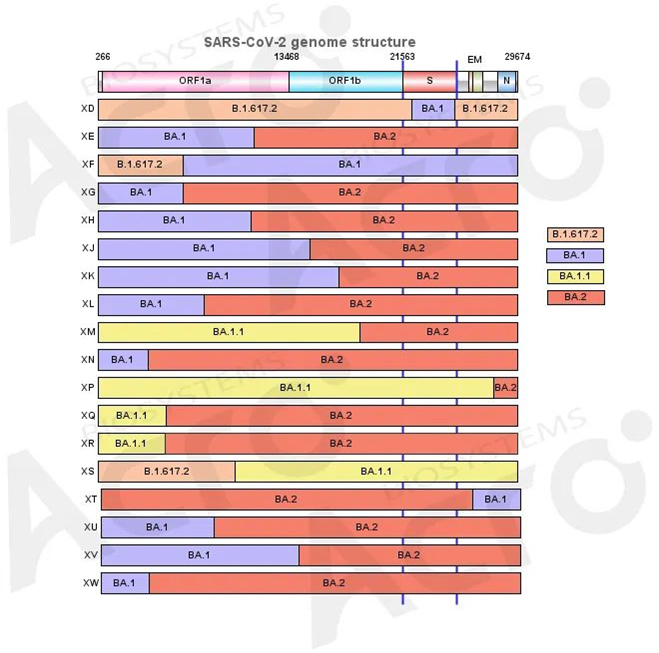
Overview of Omicron recombinant strain
Except for XD, the breakpoints in most of the recombinant strains were located in the ORF region, which means the recombination does not affect the structural protein regions, such as the spike and nucleocapsid protein. The structural proteins of these strains are exactly the same as those of the former strains, which determined that the recombination generally have no change in the recognition and antibody binding of the virus. However, since the NSP protein in the ORF region functions during viral infection, recombination in this region may affect the transmissibility of the virus.
The Omicron variant has been spreading and mutating at an alarming rate since its emergence in late 2021. The first-generation Omicron B.1.1.529 (BA.1) took over Delta's dominant position in several countries around the world in just a few weeks, until the next subvariant, BA.1.1, appeared. In March 2022, the BA.2 subvariant began to spread rapidly in parts of Asia and Europe and was considered to be more infectious than BA.1 and could escape the immune response induced by existing monoclonal antibodies. According to the statistics, BA.2 has accounted for more than 90% in the last six months. Another Omicron subtype, BA.3, was detected in South Africa at the same time but did not cause many cases of infection worldwide.
In April 2022, the number of cases of a new Omicron subvariant, BA.2.12.1, showed an explosive increase across the United States. The CDC estimates that up to early May, cases of BA.2.12.1 accounted for 43% of all infections in the United States, which is far higher then the early Febuary number which was 0.2%. Studies on the characteristics of the new variant suggest that BA.2.12.1 may be more infectious, with a transmission advantage of approximately 23% to 27% over the previously dominant (BA.2). Another sublineage of BA.2, BA.2.13, has spread in Belgium, Germany, Denmark and other European countries at the same time, infecting about 700 people.
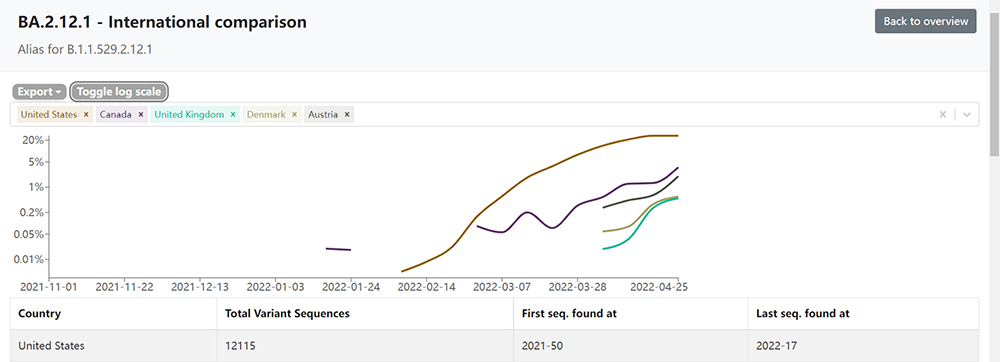
BA.2.12.1 transmission in the United States
Since the beginning of the COVID-19 outbreak, there have been successive outbreaks of several new variants in the South African region. Following Beta, Delta and Omicron (B.1.1.529), South Africa is likely to see the fifth peak of the epidemic. The World Health Organization (WHO) said that the Omicron BA.4 and BA.5 variants broke out in South Africa in January 2022 and have spread to more than 10 countries, causing more than 2000 cases.
Professor Xie Xiaoliang and his team published a pre-print article on bioRxiv on May 2, reporting their studies on the new Omicron strains, BA.2.12.1, BA.2.13, BA.4 and BA.5. The results showed that all strains above contained L452 mutations and have higher potential transmissibility than BA.2. Compared with BA.1 and BA.3, the lack of G496S mutation in BA.2 subvariants increased its ability to bind to the ACE2 receptor. However, the introduction of F486V and the loss of Q493R may lead to a decrease in hydrophobic and hydrophilic interactions, and thus significantly reduce the binding activity of BA.4 and BA.5.
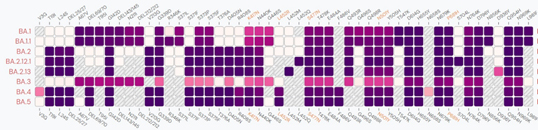
Mutation sites of Omicron lineages
Furthermore, compared with BA.2, BA.2.12.1, BA.4 and BA.5 showed stronger neutralization escape of plasma from boosted vaccines. Further analysis of the escape mutation spectrum revealed that the L452 mutation appeared under the immune stress induced by Omicron patients.
ACROBiosystems is paying close attention at the development of COVID-19. We have developed a series of new mutants' antigens at the first time, including BA.2.12.1, BA.2.13, BA.4 and BA.5. Considering the need for L452 mutations, products involving L452Q, L452M, L452R&F486V and other hot mutations were also developed. ACROBiosystems is helping to accelerate research on the efficacy of relevant mutated strains and existing vaccines, therapeutic antibodies and diagnostic reagents.
Main features:
- Covers almost all new Omicron subvariants: BA.1, BA.1.1, BA.2, BA.3, BA.2.12.1new, BA.2.13new, BA.4new, BA.5new;
- Specially developed products involve hot mutations L452Q, L452M, L452R&F486V, etc.
- Offer Spike trimer, S RBD, S1, S2, NTD and N protein in biotinylated and unconjugated versions;
- purity is more than 95% verified by SDS-PAGE and over 90% determined by MALS;
- High bioactivity verified by strict quality control, which is suitable for ELISA/SPR/BLI and other experiments.
Click to find out more information
| Lineage | Molecule | Cat. No. | Product Description |
|---|---|---|---|
| BA.2.12.1 | Spike RBD | SPD-C522q | SARS-CoV-2 Spike RBD, His Tag (BA.2.12.1/Omicron) |
| Spike protein | SPN-C522d | SARS-CoV-2 Spike Trimer, His Tag (BA.2.12.1/Omicron) | |
| BA.2.9.1/BA.2.13 | Spike RBD | SPD-C522p | SARS-CoV-2 Spike RBD, His Tag (BA.2.13/Omicron) |
| Spike protein | SPN-C5227 | SARS-CoV-2 Spike Trimer, His Tag (BA.2+L452M/Omicron) | |
| BA.3 | Spike RBD | SPD-C522i | SARS-CoV-2 Spike RBD, His Tag (BA.3/Omicron) |
| BA.4 | Spike RBD | SPD-C522r | SARS-CoV-2 Spike RBD, His Tag (BA.4&BA.5/Omicron) |
| Spike protein | SPN-C5229 | SARS-CoV-2 Spike Trimer, His Tag (BA.4/Omicron) | |
| Nucleocapsid protein | NUN-C52Hw | SARS-CoV-2 Nucleocapsid protein, His Tag (BA.4)/Omicron) | |
| BA.5 | Spike RBD | SPD-C522r | SARS-CoV-2 Spike RBD, His Tag (BA.4&BA.5/Omicron) |
| Spike protein | SPN-C522e | SARS-CoV-2 Spike Trimer, His Tag (BA.5/Omicron) | |
| Nucleocapsid protein | NUN-C52Hx | SARS-CoV-2 Nucleocapsid protein, His Tag (BA.5)/Omicron) |
Assay data
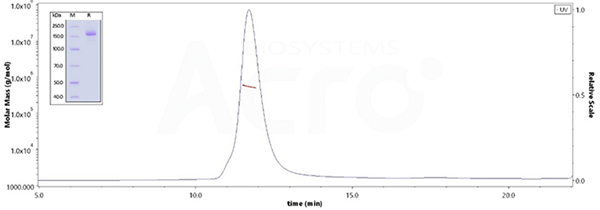
The purity of SARS-CoV-2 Spike Trimer, His Tag (BA.2.12.1/Omicron) (Cat.No. SPN-C522d) is more than 95% verified by SDS-PAGE and 90% verified by SEC-MALS. The molecular weight of this protein is around 510-560 kDa.
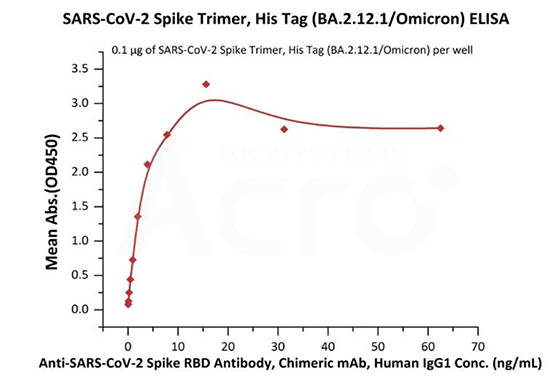
Immobilized SARS-CoV-2 Spike Trimer, His Tag (BA.2.12.1/Omicron) (Cat. No.SPN-C522d) at 1 μg/mL (100 μL/well) can bind Anti-SARS-CoV-2 Spike RBD Antibody, Chimeric mAb, Human IgG1 (Cat. No.S1N-M122)with a linear range of 0.1-4 ng/mL (Routinely tested).
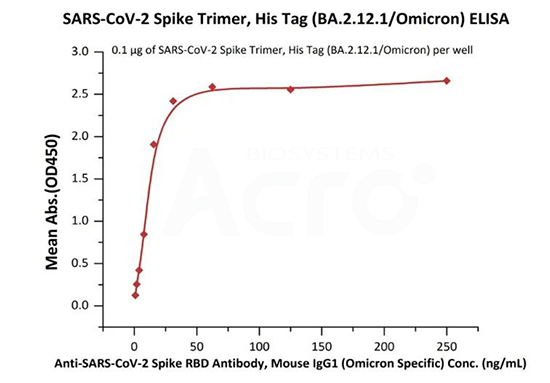
Immobilized SARS-CoV-2 Spike Trimer, His Tag (BA.2.12.1/Omicron) (Cat. No.SPN-C522d) at 1 μg/mL (100 μL/well) can bind Anti-SARS-CoV-2 Spike RBD Antibody, Mouse IgG1 (Omicron Specific) (Cat. No. SPD-M305) with a linear range of 1-31 ng/mL (Routinely tested).
1.Chen, C. et al. CoV-Spectrum: analysis of globally shared SARS-CoV-2 data to identify and characterize new variants. Bioinformatics 38, 1735–1737, doi:10.1093/bioinformatics/btab856 (2021)
2.Yunlong Cao, Ayijiang Yisimayi, Fanchong Jian, et al. BA.2.12.1, BA.4 and BA.5 escape antibodies elicited by Omicron infection. bioRxiv 2022.04.30.489997; https://doi.org/10.1101/2022.04.30.489997
This web search service is supported by Google Inc.
